About a year ago, I addressed what might seem to the average reader to be a very simple question: Why haven’t we cured cancer yet? As I pointed out at the time, it’s a question that I sometimes even ask myself, particularly given that cancer has touched my life. Three years ago, my mother-in-law died of a particularly nasty form of breast cancer. Even though I am a breast cancer surgeon, I still wonder why there was nothing that could save her (and there still is nothing that could have saved her, if it existed then) from a decline over several months followed by an unpleasant death. Yet, as a cancer researcher, I do understand somewhat. A couple of years ago, I wrote in depth about the complexity of cancer from a science-based viewpoint, as compared, of course, to the incredibly simplistic view that many purveyors of alternative medicine quackery promote as being The One True Cause of Cancer. As I put it at the time, shamelessly stealing from Douglas Adams: Cancer is complicated. You just won’t believe how vastly, hugely, mind-bogglingly complicated it is. I mean, you may think algebra is complicated, but that’s just peanuts to cancer.
I saw more evidence of that at the American Association for Cancer Research meeting last week. In fact, if there’s anything I’ve learned, it’s that developing personalized therapy for cancer is going to be a hell of a lot more difficult than we had ever suspected. Actually, it wasn’t just the AACR meeting that taught me this, but it’s as good a pretext as any to discuss some cool new science. I only wish it was science that pointed an obvious path forward to the development of personalized therapy. On the other hand, if it were easy then anyone could do the “personalized therapy for dummies” approach that, for example, Stanislaw Burzynski takes. Then there’s the even more ridiculously simplistic approach that certain practitioners of “complementary and alternative medicine” (CAM) take.
So why haven’t we cured cancer yet? Again? One reason that I discussed last time I covered this topic concerns a study that used the latest next generation sequencing (NGS) techniques to sequence seven aggressive and advanced prostate cancers. I described the results as these genomes looking like someone threw a miniature grenade into the nucleus of a prostate epithelial cell. In other words, these are some really messed up genomes. (I wanted to use another word to describe it, but this is a family blog—sort of, anyway.) I used this example to explain once again that cancer is not a single disease. It’s hundreds of diseases. Although there are common themes in how cells become cancerous, such as loss of responsiveness to growth signals with a resultant ability to grow unchecked, evasion of programmed cell death (apoptosis), inducing the surrounding tissue to provide a blood supply (angiogenesis), evading the immune system, and invading the blood or lymphatic systems to travel elsewhere in the body and take up shop in other organs, such as liver, lung, or bone, individual cancers acquire these necessary (to the cancer) abilities through many different mechanisms. For this reason, it’s completely ridiculous to speak of a “cure for cancer.”
It’s also the reason I expressed skepticism when Steve Novella discussed a potential universal anti-cancer drug. Ditto when the press breathlessly reports studies suggesting a “universal cancer vaccine.” While these sorts of research findings are promising, they need to be put into perspective. We’ve seen their like many times before, and various cancers are still deadly diseases. In fact, my career intersected with this sort of hype back in the 1990s, when I studied combining angiogenesis inhibitors with radiation therapy in experimental models of cancer in mice. For a period of time in the late 1990s, I lived the hype. Then reality, as it always does, brought us all down to earth. Now, 15 years later, we know that angiogenesis inhibitors, although useful, are not any sort of “magic bullet” cure for all solid tumors. Like many advances before, they have now taken their place in the armamentarium of anticancer drugs, more important than some but not as important as others.
It’s even more complicated than that.
The challenge
It turns out that cancer is an even more difficult foe than my original introduction of the topic indicates, which was designed to bring readers up to speed on the issue. Ever since the genomics revolution began in the 1990s and accelerated in the 2000s, the dream has been to use the gene expression profiling information and the genome sequences of individual cancers derived from next generation sequencing technology and the latest analytic techniques. It seemed like a perfectly reasonable dream early in the genomics revolution, so much so that the Cancer Genome Project, which has been sequencing cancer genomes furiously, seemed like the right way to lay the groundwork for personalized cancer therapy. Couple that with the rapid advances in sequencing technology that have allowed us to sequence both a genome and every single RNA produced by that genome, both coding and noncoding, with a rapidity previously thought completely impossible, and the rapid decline in price of sequencing a genome that might well bring the cost below $1,000 before too long, and it’s easy to understand why scientists are sequencing every tumor in sight, and more of them.
For example, a year ago a paper describing NGS sequencing of seven prostate cancers was hot news. These days, such a study would probably only make into a middle-tier journal. As of last week, one has to sequence 104 triple negative breast cancers. This was a study that was mentioned by several speakers at the AACR meeting but wasn’t published in Nature until the day after the meeting ended. This one’s so hot off the presses that it doesn’t even show up in PubMed yet. It comes from Samuel Aparicio at the University of British Columbia, plus other groups from UCSF, Cambridge, the University of Alberta, and Simon Fraser University and is entitled The clonal and mutational evolution spectrum of primary triple-negative breast cancers. A news report describes the finding thusly:
The journal Nature has just published the team’s findings online.
The study is the largest genetic analysis of what were thought to be triple negative breast cancer tumours.
The 59 scientists involved in this study expected to see similar gene profiles when they mapped on computer the genomes of 100 tumours.
But to their amazement they found no two genomes were similar, never mind the same. “Seeing these tumours at a molecular level has taught us we’re dealing with a continuum of different types of breast cancer here, not just one,” explains Steven Jones, co-author of this study.
Triple-negative breast cancer (TNBC) is a form of breast cancer that is characterized in the clinic by its lack of hormone receptors—i.e., estrogen receptor (ER) and progesterone receptor (PR)—and its lack of amplification of HER2/neu, an oncogene that can be targeted with Herceptin (trastuzumab). In general, most (but not all) TNBCs belong to a category of breast cancer established 10 years ago by early gene expression profiling techniques known as basal-like. Worse, TNBC tends to be more aggressive than ER-positive cancers, metastasizing earlier and recurring earlier. Because TNBC doesn’t express receptors that confer sensitivity to agents targeted against the estrogen receptor or HER2, the only systemic therapy that is currently effective against TNBC is cytotoxic chemotherapy, which is very effective initially. Unfortunately, that efficacy doesn’t translate into better survival. TNBC, although making up only around 15% of breast cancer, accounts for a disproportionate share of breast cancer deaths. This study, which looked at newly diagnosed TNBC before any treatment was initiated, suggests one reason why. In this study of 104 TNBCs, which is the largest series of tumors subjected to NGS analyses to date, no two tumors were the same, and they were heterogeneous even before any treatment was initiated.
That reason is that TNBC is not a single disease. In fact, even an individual TNBC tumor is not a single disease. Tumor cells undergo evolution as they grow, so that the cells in them are genetically heterogeneous. The cells growing in one area of a tumor can and often do have markedly different genetic mutations from the cells growing in another part of the tumor. Arapicio’s group found evidence that this is going on in TNBC in spades. One finding in the study was that groups of mutations within individual cases have different clonal frequencies, indicative of distinct clonal genotypes. What this means is that analysis of the NGS data found evidence of different clones of cells with different frequencies of groups of mutations, suggesting that there are a number of different clones of cells with different genetic makeups in each tumor. They also found that p53 was the most frequently mutated gene and that tumors harboring p53 mutations tended to have the greatest number of overall mutations, consistent with the role of p53 as “guardian of the genome.” Moreover, tumors varied wildly from each other in the number of mutations, with some tumors harboring just a handful of somatic mutations and others harboring hundreds. Finally, only around 36% of these mutations were expressed, which is consistent with other studies. Overall, this study shows:
The team found that each tumor displayed multiple “clonal genotypes,” suggesting that the cancer would have to be treated as multiple diseases, rather than a single entity.
“Triple negative breast cancer is not just one uniform subtype of breast cancer,” Sam Aparicio, professor of pathology and lab medicine at UBC and senior author of the study, said in a statement. “It’s actually extremely complex, with each cancer at a different stage in the evolutionary process at the time of diagnosis.” This finding could “help explain why patient responses to treatment differ greatly,” he added.
If there’s a place where evolution will inform medicine at least as much as it does in infectious disease and the development of antibiotic resistance in microorganisms, it’s in cancer. The knowledge that tumors are heterogeneous and that they undergo evolution based on the selective pressures from the host, including characteristics of the organ and its microenvironment and other factors, is not new. Neither is the knowledge that cancer cells undergo evolution under the selection pressure of various therapies, thus developing resistance to those therapies. Indeed, it is the tumors that are less heterogeneous because they are driven by a single mutation, that tend to be actually curable or best controlled with single-agent targeted therapies. Just like the case in whole organisms undergoing evolution in response to selective pressures, genetic variability in cancer cells provides the raw material upon which evolutionary selection pressures can operate. The result is that tumors with more genetic heterogeneity tend to develop resistance to therapy sooner.
Personalized therapy versus cancer cell heterogeneity
Less than a month before the AACR meeting, yet another study was published that threw cold water on the idea that NGS sequencing of tumors will lead to personalized therapy of cancer any time soon. This study by Charles Swanton and colleagues published in the New England Journal of Medicine, is yet more evidence at how heterogeneous cancer is and how it evolves as it progresses and metastasizes. Basically, what Swanton’s group did was to examine four patients with metastatic renal cell carcinoma, but to analyze different locations within the primary tumor and metastases, subjecting the tissue from these various locations to NGS techniques and mutational analysis.
Data derived from the first patient is probably the best example to discuss. This is a patient who had a type of renal cell carcinoma known as clear cell carcinoma, with lung and chest wall metastases. He was treated for six weeks with everolimus and then underwent nephrectomy. He was started on everolimus again for six weeks and then underwent resection of his chest wall metastases. In both cases, the tumors did not shrink in response to the targeted treatment. Investigators then did the following:
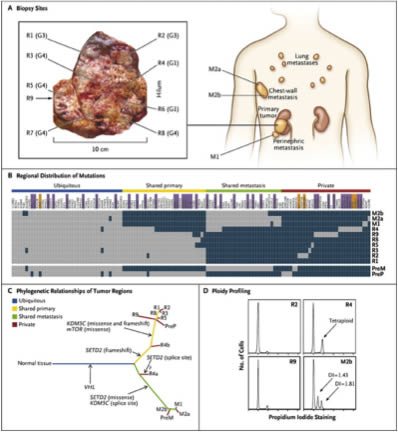
For Patient 1, we performed exon-capture multiregion sequencing on DNA from pretreatment biopsy samples of the primary tumor (PreP) and chest-wall metastasis (PreM), nine primary-tumor regions of the nephrectomy specimen (R1 to R9), a metastasis in the perinephric fat of the nephrectomy specimen (M1), two regions of the excised chest-wall metastasis (M2a and M2b), and germline DNA19 (Figure 2A). This sequencing resulted in a median coverage of 74 reads (Table 1 in the Supplementary Appendix). Nonsynonymous somatic point mutations and insertions and deletions (indels) that change the protein amino acid sequence were filtered and manually reviewed to remove sequencing and alignment errors and to determine the regional distribution of mutations. Regions R6 and R7 were excluded from analyses since only one nonsynonymous variant passed filtering. We identified 101 nonsynonymous point mutations and 32 indels (Table 2 in the Supplementary Appendix) and mapped their regional distributions across the tumor (Figure 2B). Sanger sequencing was used to validate 42 mutations. Of these mutations, 37 (88%) were specifically validated in regions in which they were initially identified (Figure 2B, and Fig. 1 in the Supplementary Appendix), documenting genetic intratumor heterogeneity.
Don’t worry so much about the gobbledygook. The point is that the investigators sequenced nine areas in the primary tumor, two regions of the chest wall metastasis, a metastasis in the fat around the kidney, and the patient’s germline DNA for comparison. What they found was a pattern of mutations in the tumor regions consistent with branching evolution:
We inferred ancestral relationships and constructed a phylogenetic tree of the tumor regions by clonal ordering, as described by Merlo et al.20 (Figure 2C), which revealed branching rather than linear tumor evolution. One branch evolved into the clones present in metastatic sites, and the other diversified into primary tumor regions. R4 shared some, but not all, primary-tumor and metastatic mutations, which suggested the presence of at least two clonal populations in this region that arose from progenitor cells of the metastases and of other primary tumor sites.
And, since a picture is worth a thousand words, here is Figure 2:
Another key finding is that it really matters which part of a tumor a biopsy is taken from. It matters a lot. For instance, in this same tumor, a single biopsy only contained approximately 55% of all the mutations detected in the tumor and only 34% of all mutations that were detected by multiregion sequencing in the nephrectomy specimen were present in all regions. To me, it’s a classic scientific-ese understatement when the authors state that these results indicate “that a single biopsy was not representative of the mutational landscape of the entire tumor bulk.”
This heterogeneity means something, too. More and more, oncologists are using tests that are gene expression signatures in order to guide therapy. In breast cancer, for instance, we use the Oncotype DX assay a lot in order to decide whether patients with ER-positive breast cancers and no lymph node metastases will benefit from chemotherapy. In renal cell carcinoma, apparently there is a 110-gene signature profile that is used to classify clear cell carcinoma. Swanton looked specifically at this profile in Patient #1:
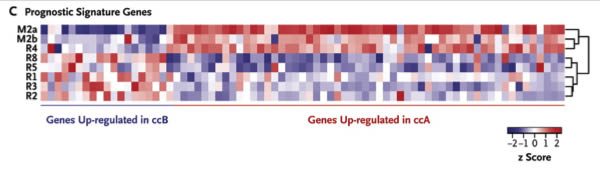
We determined the intratumoral expression of a 110-gene signature shown to classify clear-cell carcinoma into two molecular subgroups: clear-cell A (associated with a good prognosis) and clear-cell B (associated with a poor prognosis).24 Consistent with the phylogenetic analysis, metastatic sites and the primary-tumor site R4 segregated together, enriched for genes in the clear-cell A subgroup, in contrast to the remaining tumor regions that were enriched for the clear-cell B subgroup (Figure 3C). Thus, prognostic gene-expression signatures may not correctly predict outcomes if they are assessed from a single region of a heterogeneous tumor.
To give you a visual picture, take a look at Figure 3C. You don’t need to know how to interpret the gene signatures, other than that red means the gene is expressed at a higher level and blue means it’s expressed at a lower level. Just look at the patterns in the different areas of the tumor assayed. Notice how different they are? That’s what the authors are talking about. Once again, a picture is worth the proverbial thousand words (not that that stops me from writing a thousand words anyway).
Given the difficulty we often have in obtaining anything more than a needle biopsy of tumors, one biopsy from one area of a tumor is almost always what oncologists base their treatment decisions on. At least in breast cancer, we usually have the whole primary tumor, but even so only a small portion is used to do the actual Oncotype DX test. What are the consequences of this tumor heterogeneity? Evolution rules. For instance, a few months ago, researchers at Washington University did NGS analyses on the genomes of primary and relapsed tumors in eight patients with acute myeloid leukemia (AML) and found two main patterns in the relapsed tumors:
In addition to discovering novel, recurrently mutated genes (for example, WAC, SMC3, DIS3, DDX41 and DAXX) in AML, we also found two major clonal evolution patterns during AML relapse: (1) the founding clone in the primary tumour gained mutations and evolved into the relapse clone, or (2) a subclone of the founding clone survived initial therapy, gained additional mutations and expanded at relapse. In all cases, chemotherapy failed to eradicate the founding clone. The comparison of relapse-specific versus primary tumour mutations in all eight cases revealed an increase in transversions, probably due to DNA damage caused by cytotoxic chemotherapy. These data demonstrate that AML relapse is associated with the addition of new mutations and clonal evolution, which is shaped, in part, by the chemotherapy that the patients receive to establish and maintain remissions.
Inside Darwin’s tumor, indeed. The tumor cells that survived chemotherapy acquired new mutations that allowed them to regrow even faster. The parallels between cancer biology and antibiotic resistance are both sobering and humbling, the attempts of evolution denialists to paint evolution as irrelevant to medicine notwithstanding.
Meanwhile, at the AACR meeting, there were a number of other abstracts and talks that were variations on the same theme. There is a large degree of inter-tumor variability in genetic makeup and a large degree of intra-tumor variability as well. One of these studies, which looked at tumors from the ACOSOG Z1031 trial comparing different aromatase inhibitors in ER-positive breast cancer, found that the complexity of genomic changes correlated with lack of response to therapy. Moreover, as was discussed in at least a couple of talks at the AACR meeting, an evolution-based analysis of tumors using the latest NGS techniques indicates that for most solid tumors the time from founder mutation to clinically apparent metastasis is between 20 and 30 years. For example, for pancreatic cancer it’s a median of around 21 years, and for colon cancer it’s around 30 years. That’s decades of evolution that can occur in the tumor to result in heterogeneity. Worse, it means that, by the time we see a tumor clinically, it’s already very late in the process. This is not new news, of course. We’ve always known that it’s at least several years from initiation of cancer to clinical presentation, with the part of the process from detection of the tumor to death making up a relatively small part of the timeline of the history of a cancer. All that the new NGS sequencing techniques have shown us is that the process of carcinogenesis starts earlier than we had previously thought and results in more branches in the evolutionary tree of a tumor than we had previously thought, resulting in even more tumor heterogeneity than we had thought.
So what the new genomic analytical techniques have re-emphasized to us in a quantitative manner that we couldn’t use before is that not only is cancer not a single disease but that cancer types are not single diseases. Breast cancers, for instance, have been shown to harbor at least 1,700 different mutations, but only three of them showed up in at least 10% of patients, with the great majority of them being unique to each patient. The most recent study in Nature discussed above shows that even a single subtype of breast cancer that is generally treated clinically more or less the same, TNBC, varies wildly (and almost continuously) in the genomic changes and mutations each tumor has. Not only that, but each TNBC is in essence several diseases, because each TNBC is made up of many different clones that have evolved as the tumor itself grew, progressed, and evolved. All of this occurs even before the tumor has been subjected to any treatment at all. As if that’s not bad enough, it would appear that tumors are a mosaic of groups of many different tumor cell types that develop through branching evolution such that metastases can be very different from the primary tumor and even different regions of the primary tumor can be very different from each other, so much so that finding a “favorable prognosis signature” on a core biopsy means only that that one area biopsied has that gene signature. Large areas elsewhere in the tumor could have the unfavorable prognosis signature.
In an accompanying editorial in the NEJM, Dan DeLongo sums it up thusly, puncturing the optimism at the heart of visions of personalized medicine for cancer using targeting agents:
This modicum of success (since none of the new agents produce cures in advanced cancers, although curative combinations may yet emerge) has led to a certain level of overoptimism in the field. A new world has been anticipated in which patients will undergo a needle biopsy of a tumor in the outpatient clinic, and a little while later, an active treatment will be devised for each patient on the basis of the distinctive genetic characteristics of the tumor. The path to that new world is already being cleared, with several companies now marketing genetic tests that measure the genetic signature of a tumor, with the expectation that this signature will direct the choice of treatment and predict treatment outcome.
However, as Albert Einstein noted, “things should be made as simple as possible, but not simpler.” A serious flaw in the imagined future of oncology is its underestimation of tumor heterogeneity — not just heterogeneity between tumors, which is a central feature of the new image of personalized medicine, but heterogeneity within an individual tumor.
Derek Lowe put it quite well in the title of a recent post, Personalized Medicine for Cancer? Try Every Cell. He’s right. We’re at the point where it is now possible to do NGS on single cells. If the rate of progress continues apace, a year or two from now such sequencing studies will become fairly routine. At least at the big genomics centers, it’s becoming more common. As these studies go from sequencing 100 cells to sequencing 1,000 cells or 10,000 cells, or a million cells, it’s likely that it will be discovered that intratumor heterogeneity is even greater than found in these two studies.
The future of personalized medicine
Does all of this mean that personalized cancer therapy is a pipe dream? If I thought that, I wouldn’t be moving towards genomics in my research. It is, however, becoming clear that developing personalized therapies for cancer is going to be a lot more difficult than we had thought before. I asked the rhetorical question, “Why haven’t we cured cancer yet?” The answer is, of course, that curing a single cancer, much less many cancers, is really, really hard. Scientists not only have to contend with basically a continuous spectrum of mutations between individual cancers that renders the concept of even single cancer subtypes horribly naive, but with a near continuous spectrum of mutations between groups of cells in a single tumor.
There is reason for hope, however. As was discussed at AACR, the vast majority of cancer-causing driver mutations can be divided into 12 key molecular signaling pathways, and, as was found in the NEJM paper, mutations at the “trunk” of the divergent branching evolution tend to remain as divergent evolution occurs, and targeting them is likely to be more effective than targeting mutations further out on the branches. It might be possible to hone in on these different pathways. In the meantime, as the cost of sequencing continues to fall, it might become feasible to sequence several parts of a tumor and its metastases and map out treatments that cover all of them. Personalized medicine is thus possible and still holds promise. It’s just not going to be as simple as getting a biopsy of a tumor and picking a targeted agent or two to blast the tumor into oblivion. The point is not that personalized medicine is impossible or that it’s impossible to develop cures for various cancers. The point is that biology is always way more complicated than we ever thought it was, and evolution almost always wins out. Or, as a character named Dr. Ian Malcolm in the movie Jurassic Park, observes:
If there is one thing the history of evolution has taught us it’s that life will not be contained. Life breaks free, expands to new territories, and crashes through barriers, painfully, maybe even dangerously, but, ah, well, there it is.
Unfortunately, paraphrase a later quote from Dr. Malcolm, cancer always finds a way. Well, pretty darned close to always, anyway. If I believed that cancer always finds a way, then I would also believe that cancer research and personalized therapy are futile endeavors. I do not believe that.
Even so, the reason we haven’t cured cancer yet is because we haven’t figured out how to overcome the power of evolution. Until we figure out a way to do that, we will continue to make only incremental progress.